Simple and sustainable R packaging using inlinedocs
You wrote a bunch of R code, now you want to share it with
everyone, so you are making a package. Most likely you have a bunch of
functions in a file, and now you have to write all the documentation
for them. Now how to go about doing that?
- Option 1: do it yourself. Fire up package.skeleton() and
then edit the Rd files by hand, following the directions in Writing R
Extensions (over 100 pages in PDF form) --- good luck. It takes a
long time to edit all these files by hand, and if you change your code
you will end up with docs which do not agree. Unless you constantly
update all those Rd files. (will you really?)
- Option 2: put documentation in big header comments, as
in
roxygen,
roxygen2,
mvbutils::doc2Rd,
or
R.oo::Rdoc$compile.
You like the idea of combining source code with documentation, because
then you can do "literate programming" and moreover, it is practically
easier to keep them in sync with each other. But these packages have a
number of drawbacks, especially for documenting function
arguments. The huge headers are usually far away from the actual code
being documented, and you have to repeat yourself when you name the
function arguments in the comment and the definition. (but to its
credit, the call graphs that roxygen makes are pretty cool).
- Option 3: inlinedocs. You will never have to repeat
yourself, just comment your code in natural places, and you get Rd
files that pass R CMD check so you can publish that package and move
on with your life. And when you make changes to that function, the
docs are right next to the code so it is easy to update the docs.
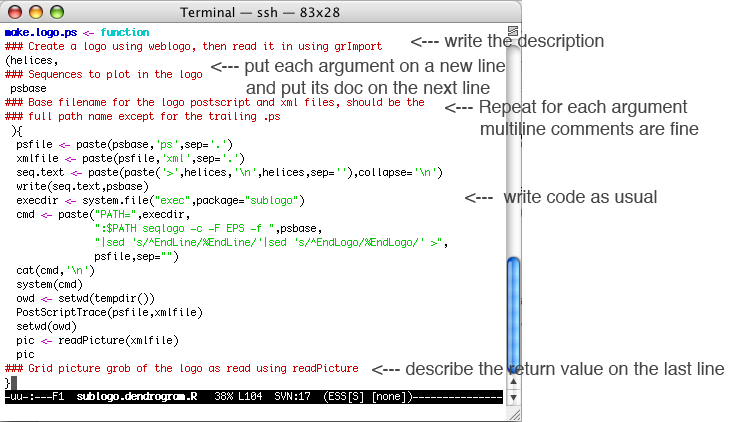
Then put your code in pkgdir/R/code.file.R, write a pkgdir/DESCRIPTION,
start R, and do:
## R-Forge and CRAN versions of inlinedocs are deprecated since 3 June 2014.
## Instead, install from the updated version on GitHub:
install.packages("devtools")
devtools::install_github("tdhock/inlinedocs")
library(inlinedocs)
package.skeleton.dx("/path/to/your/pkgdir")
Then you are done! You will get Rd files in pkgdir/man that are
guaranteed to pass R CMD check! Some notes/tips:
- Use an editor where you can easily do line breaks in your
comments, like Emacs
with ESS. Then you just type
### and go on as long as you want with your comment. When
you're done and you have a really long comment, do a M-q to
automatically break lines and add ### prefixes.
- You can add a description of a data item declared in your code
by
adding a ### comment on the line before the variable is first
declared.
- By default, R code in *.r files will not be used for inlinedocs.
If you want to write some *.Rd files by hand then you do not want
inlinedocs to overwrite those files, so put that code in *.r files. If
you want inlinedocs to process *.r files, then you can specify
package.skeleton.dx(excludePattern=NULL).
- The DESCRIPTION file is used by inlinedocs, so please
fill in at least these fields: Package Version License Description
Title Author Maintainer. The most important field
is Author, which is copied to the author section of every
generated Rd file. If you haven't written one
yet, package.skeleton.dx() will create an
empty DESCRIPTION file for you.
- You still have to document your datasets (the files in
pkgdir/data) by writing an Rd file, but these don't change
often, so that's no problem. To get started just use
the prompt() function.
- To add inline examples for FUN, use the ex
attribute of FUN, as shown below. This is more sustainable
than putting test code in R comments, since debugging and changing
commented code is a pain.
- For additional flexibility, inlinedocs now allows documentation of
S4 class definitions (i.e. those which use
setClass) and includes additional triggers based on the
string ##<<. This means that most of the standard
documentation sections can be filled in from wherever is relevant in
the source code, as follows:
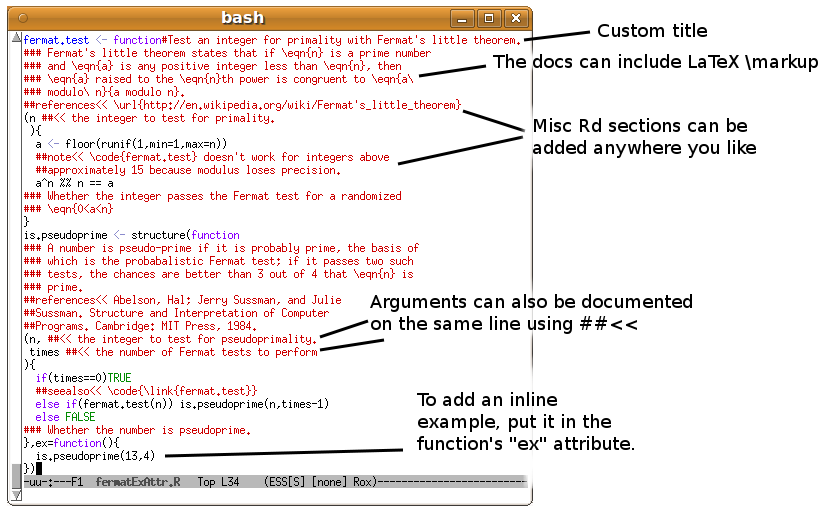

- ##<< at the end of a line can be used to add
function argument descriptions on the same line as an argument name,
and
any
immediately following ## comment lines will also be
included.
- ##xxx<< as the first non-white space element on a
line introduces a section of consecutive ## comment lines
which will be placed in the xxx section of the documentation,
where xxx is one of:
alias,
details,
keyword,
references,
author,
note,
seealso,
value,
title or
description. Multiple such chunks are concatenated
appropriately, so you can think of this as something like the C++
"put" operator putting chunks into that section. For further details
and examples of these look for << in the source
file parsers.R.
- ##describe<< within such a documentation chunk
allows a "describe" block within that chunk to be constructed using
same-line ##<< comments (as for the function
arguments). All the following source lines until
another ##xxx<< line will be scanned for such same-line
comments. ##end<< may be used to end such a block and
return to the same documentation chunk. This
allows named
lists to be documented using the names actually in the code so
that it is less easy to forget to document when you add an
element.
- Be careful to use the right number of # ...
- # for the title.
- ##<< for documenting function arguments on the
same line.
- ##section<< followed by ## for arbitrary
documentation sections.
- ### for description, function arguments, and return
values.
-
Some more examples of inlinedocs syntax can be found in the
testfiles
directory. These files contain the unit tests which are checked
before publishing each new version of inlinedocs, so you can be sure
that they will work correctly.
Extending the inlinedocs syntax with your own Parser Functions
inlinedocs operates using a modular system of source code parsers,
so it is very simple to extend. To do this, you just need to write a
custom Parser Function that takes your source as input and returns a
list of documentation objects. For example, see the simple
function in
testfiles/alternate.R,
which can be used as a Parser Function in the parsers
argument to package.skeleton.dx().
Patch for ESS
The useful C-c C-f ESS command currently doesn't work with some valid
R inlinedocs function definitions. A temporary remedy is to download
the ESS source and change the ess-function-pattern variable:
wget http://ess.r-project.org/downloads/ess/ess-5.13.tgz
tar -xzf ess-5.13.tgz
cd ess-5.13
wget https://r-forge.r-project.org/scm/viewvc.php/*checkout*/lisp/ess-function-pattern-patch/ess-function-pattern-more-general.patch?root=inlinedocs -O ess-function-pattern.patch
patch -p0 < ess-function-pattern.patch
Return to
the project
summary page.


